Credit: Ernesto del Aguila III, National Human Genome Research Institute, NIH
Using weakly electric fish to understand the evolution of convergently evolved, novel phenotypic traits
Jason Gallant
Michigan State University, USA
Electric fish have served as a model system in biology since the 18th century, providing insight into the nature of bioelectrogenesis, the structure of the synapse, and brain circuitry underlying complex behavior. Electric fishes exhibit many interesting phenotypes, but very little is known about how they are encoded in the genome, which is a broad goal in 21st century biology. Electric fish offer a singular advantage in trying to understand the link between the genome and the phenotype it produces: two groups of electric fish evolved independently in Africa and South America, providing a system where neuroscientists, evolutionary biologists, developmental biologists and cell biologists could examine the genetic mechanisms of convergently evolved phenotypes with built-in replication.
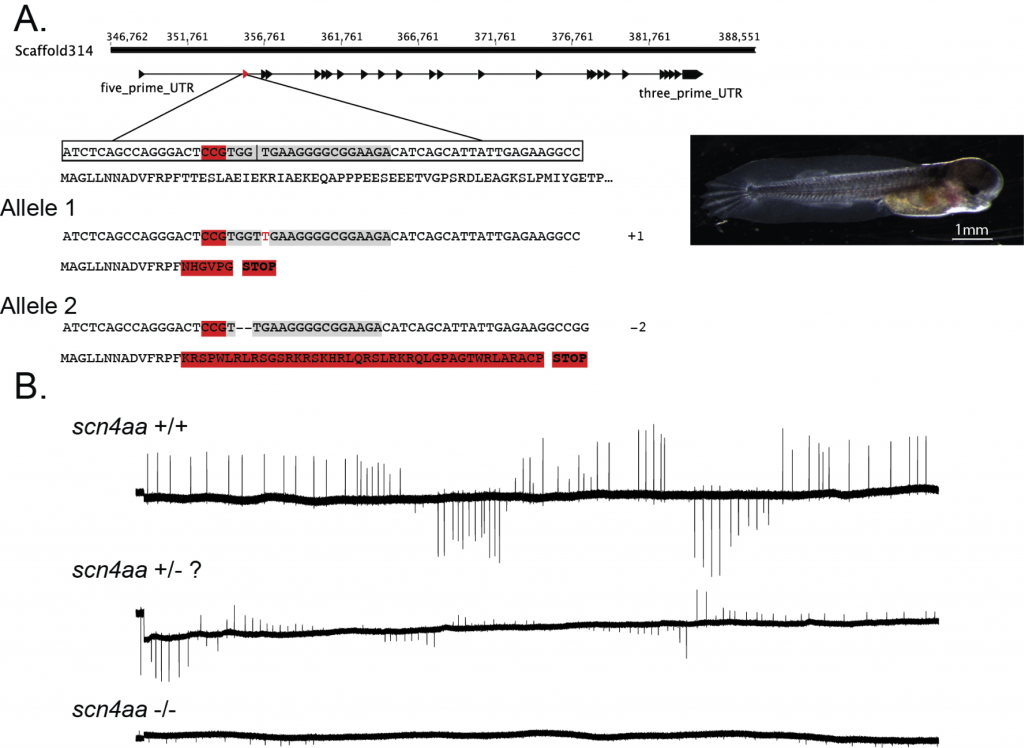
While there is a growing amount of electric fish genomic data, there are no functional tools to investigate this connection. I will report on ongoing efforts among a network of US biologists to develop a robust, accessible, and easily transferable genetic manipulation toolbox for the electric fish model system. The first phase of the project is rapid parallel protocol development for CRIPSR/Cas9 gene editing, the use of vivo-morpholinos, and viral-mediated gene transfer approaches in electric fish. The second phase of the project will be to disseminate protocols and datasets by developing web-based software for sharing protocols, data, and resources, and to train the next generation of electric fish biologists to use these techniques.
This work is supported by NSF #1644965 to J.R.G.


You must be logged in to post a comment.